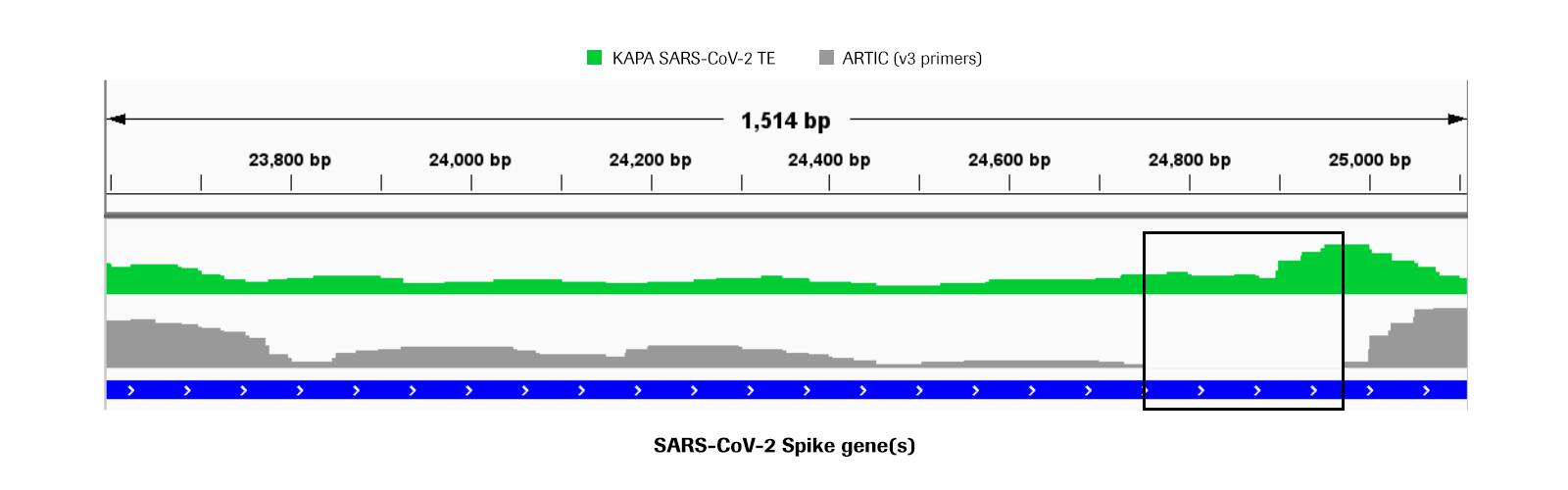
Figure 2. KAPA SARS-CoV-2 panel achieves more uniform coverage compared to the ARTIC amplicon method. IGV coverage plot of one representative replicate of the KAPA SARS-CoV-2 TE dataset (green) compared to a replicate from the ARTIC workflow with v3 primers (gray). A near dropout of reads (black box) was observed in the Spike gene for the ARTIC libraries in all 3 replicates tested while the KAPA SARS-CoV-2 TE yielded more even coverage across the whole Spike region.