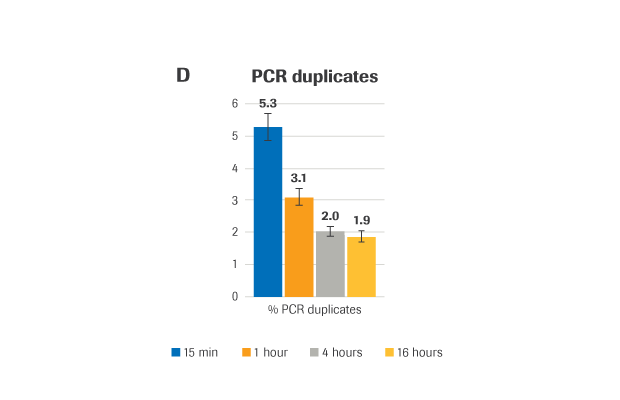
KAPA HyperExome yields high-quality results with hybridization times as short as 1 hour
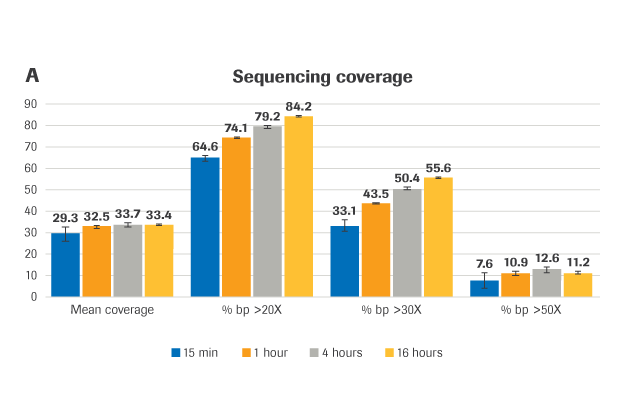
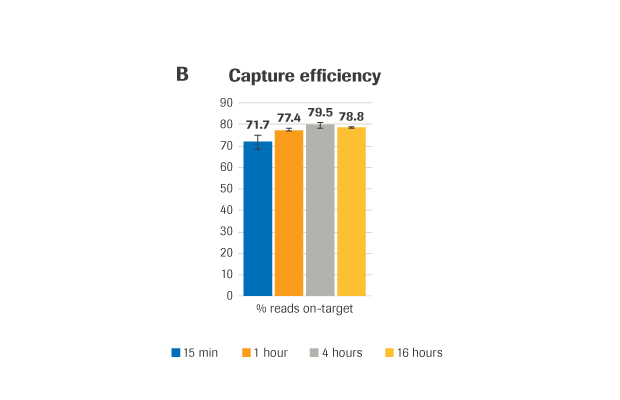
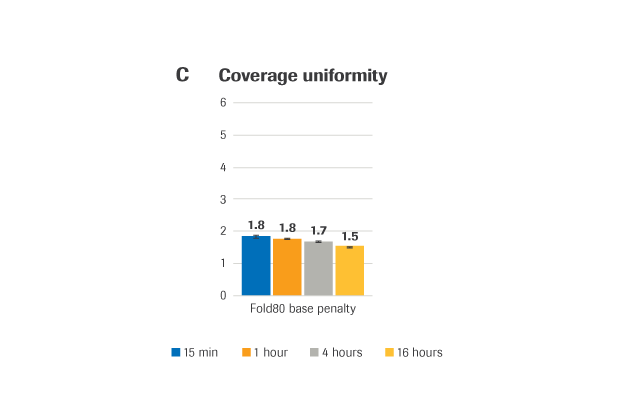
(A) Sequencing coverage (B) Capture efficiency, presented as % reads on-target (the percent of mapped, non-duplicate reads overlapping the target region by at least 1 base). (C) Coverage uniformity, presented as Fold-80 base penalty. (D) PCR duplicates, a measure of library complexity (fewer % PCR duplicates=greater library complexity).
METHOD: Target-enriched libraries were generated using the HyperCap Workflow v3.0, with the KAPA HyperPrep Kit and KAPA HyperExome Probes. Single-plex hybridization reactions were carried out at 55°C with 1 μg library and KAPA HyperExome probes, for the following durations: 15 minutes, 1 hour, 4 hours, and 16 hours (standard hybridization is 16 – 20 hours). Normalized, pooled libraries were sequenced on an Illumina NextSeq 500 instrument using the NextSeq High Output kit (2 x 75 bp). Data was down-sampled to 50X raw coverage. For all charts, bars represent the mean from triplicate libraries and error bars indicate the standard deviation. Note: this protocol is still in development and has not yet been fully validated.